15.3: Constructing Network Models with NetworkX
- Page ID
- 7859
Now that we have finished the above crash course on graph theoretic terminologies, it is time to begin with computational modeling of networks. As briefly previewed in Sections 5.4 and 12.2, there is a wonderful Python module called NetworkX [27] for network modeling and analysis. It is a free network analysis toolkit widely used by network researchers. If you are using Anaconda, NetworkX is already installed. If you are using Enthought Canopy, you can still easily install NetworkX by using its Package Manager. You can find the documentation for NetworkX online at http://networkx.github.io.
Perhaps it is worth mentioning why we are choosing NetworkX over other options. First, its functions are all written in Python, and their source codes are all available on the website above, so you can check the details of their algorithms and even modify the codes if you want. Second, NetworkX uses Python’s plain “dictionaries” to store information about networks, which are fully accessible and flexible to store additional information you want to add. This property is particularly important when we implement simulation models of dynamical networks. Specialized data structures implemented in other network analysis tools may not have this flexibility.
So, let’s first talk about the data structure used in NetworkX. There are the following four different data types (called “classes”) that NetworkX offers:
Graph For undirected simple graphs (self-loops are allowed)
DiGraph For directed simple graphs (self-loops are allowed)
MultiGraph For undirected multigraphs (self-loops and multiple edges are allowed)
MultiDiGraph For directed multigraphs (self-loops and multiple edges are allowed)
You can choose one of these four data types that suits your modeling purposes. In this textbook, we will use Graph and DiGraph mostly.
You can construct a graph of your own manually. Here is an example:
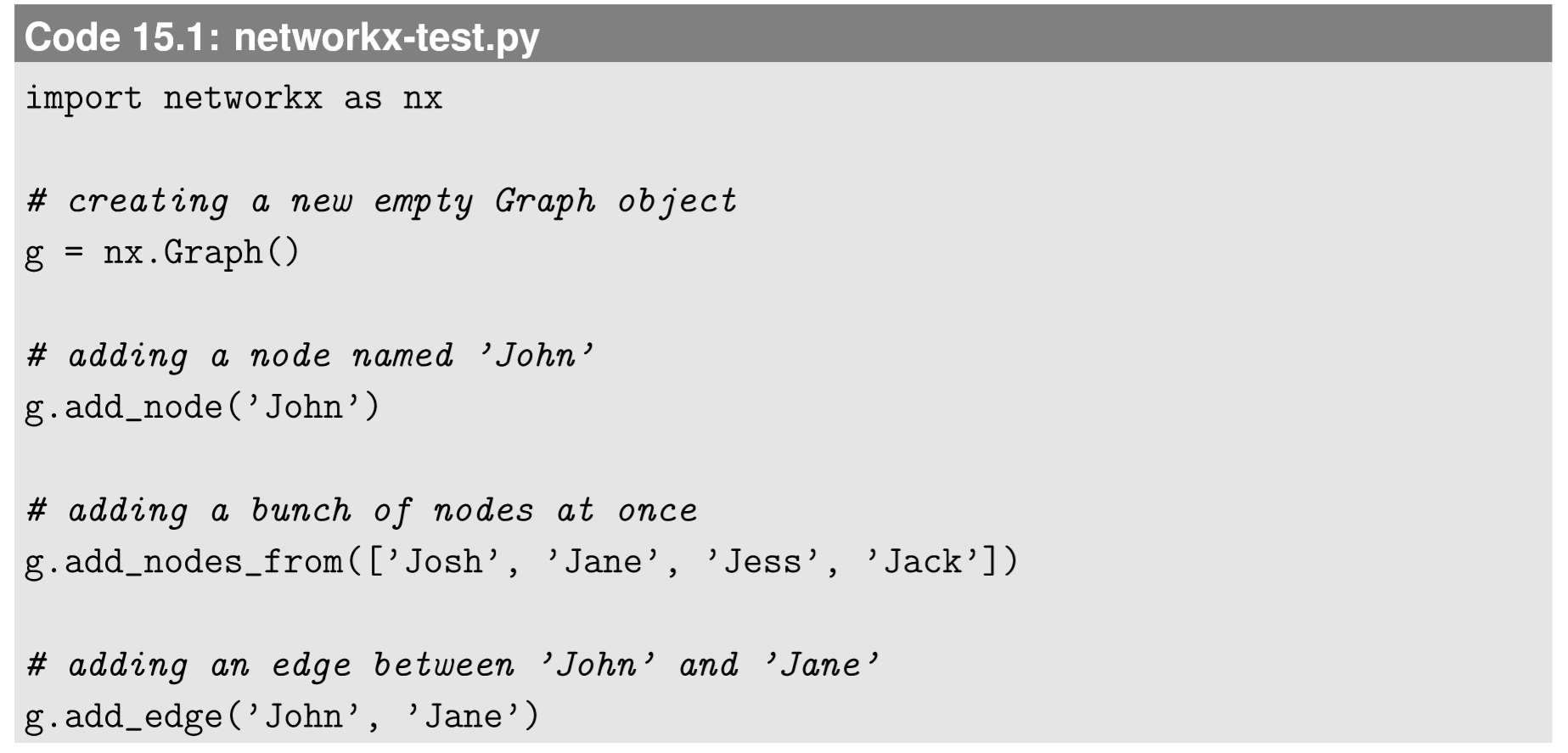


Here I used strings for the names of the nodes, but a node’s name can be a number, a string, a list, or any “hashable” object in Python. For example, in the phase space visualization code in Section 5.4, we used a tuple for a name of each node.
I believe each command used above to add or remove nodes/edges is quite self-explanatory. If not, you can always look at NetworkX’s online documentation to learn more about how each command works. There are also several other ways to manipulate a graph object, which are not detailed here.
Once this code is executed, you can see the results in the Python command line, like this:

The first output (“<networkx.classes. ... >”) shows that g is a Graph object. In order to see the contents of the data in the object, we need to use some commands. g.nodes() returns a list of all nodes in the network, while g.edges() returns a list of all edges. Compare these results with the commands we used above to check if the node/edge additions/removals were conducted correctly.
Those nodes() and edges() are functions that read the raw data inside the Graph object g and then produce a cleaned-up list of nodes or edges. But NetworkX also allows you to have direct access to the Graph object’s internal raw data structure. The data about nodes are stored in a dictionary called node right under g:

This is a dictionary whose keys and values are the nodes’ names and their properties, respectively. The properties of each node are also stored in a dictionary (initially they are all empty, as shown above), so you can dynamically add or modify any node property as follows:

We will use this method to add dynamic states to the nodes in Section 16.2.
Similarly, the data about the edges are also stored in a dictionary called edge under g:

This is a little hard to read, so let me insert line breaks at the appropriate places so that the structure of the data is clearer:
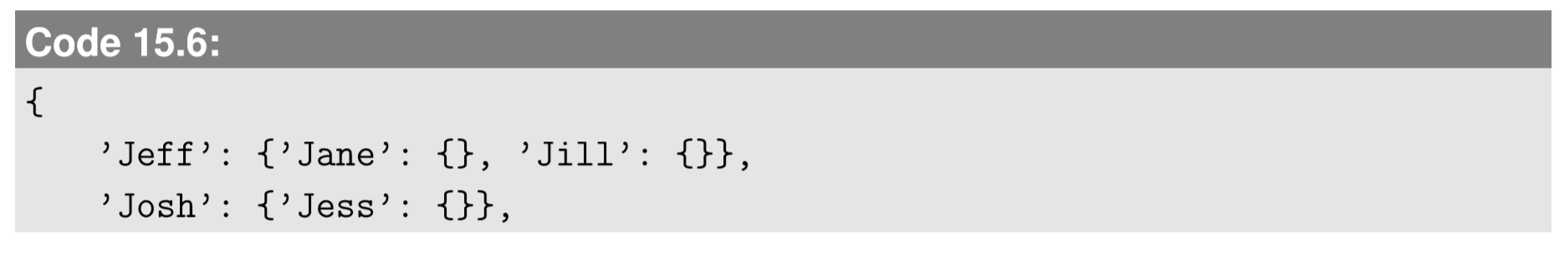

Now its structure is much clearer. g.edge is a dictionary that describes edges in the network in the adjacency list format. Each entry of g.edge is a pair of the node’s name and a dictionary that contains its neighbors. For example, the first entry says that Jeff is connected to Jane and Jill. Note that the connections are perfectly symmetric; if Jeff is connected to Jane, then the entry for Jane (in the last line) also contains Jeff as her neighbor. Such symmetry is automatically maintained by NetworkX, because this Graph object is for undirected graphs.
Moreover, each neighbor’s name works as a key associated with another dictionary in which we can store properties of the connection (initially they are all empty, as shown above). So you can dynamically add or modify any edge property as follows:

Again, the output is reformatted to enhance the readability. Note that the new edge properties (’trust’ between Jeff and Jane; ’love’ between Josh and Jess) are correctly inserted into the respective dictionaries, always maintaining symmetry between nodes. I just added True ’love’ from Josh to Jess, which has been reciprocated from Jess to Josh! What a beautiful world. This is all because we are using a Graph object.
The DiGraph object behaves differently. It is designed to represent directed graphs, so the edges are all considered one-way. Symmetries are not maintained automatically, as illustrated in the following example:

The last error message means that Jess doesn’t know a guy named Josh in this case, because the graph is asymmetric. Life is hard.
Represent the network shown in Exercise 15.1 as a Graph object of NetworkX.
Represent a small social network around you (say, 10 people) as either Graph or DiGraph object of NetworkX. Then add properties to nodes and edges, such as:
• Node properties: full name, age, job, address, etc.
• Edge properties: relationship, connection weight, etc.
In the examples above, we manually constructed network models by adding or removing nodes and edges. But NetworkX also has some built-in functions that can generate networks of specific shapes more easily. Here are a few examples:
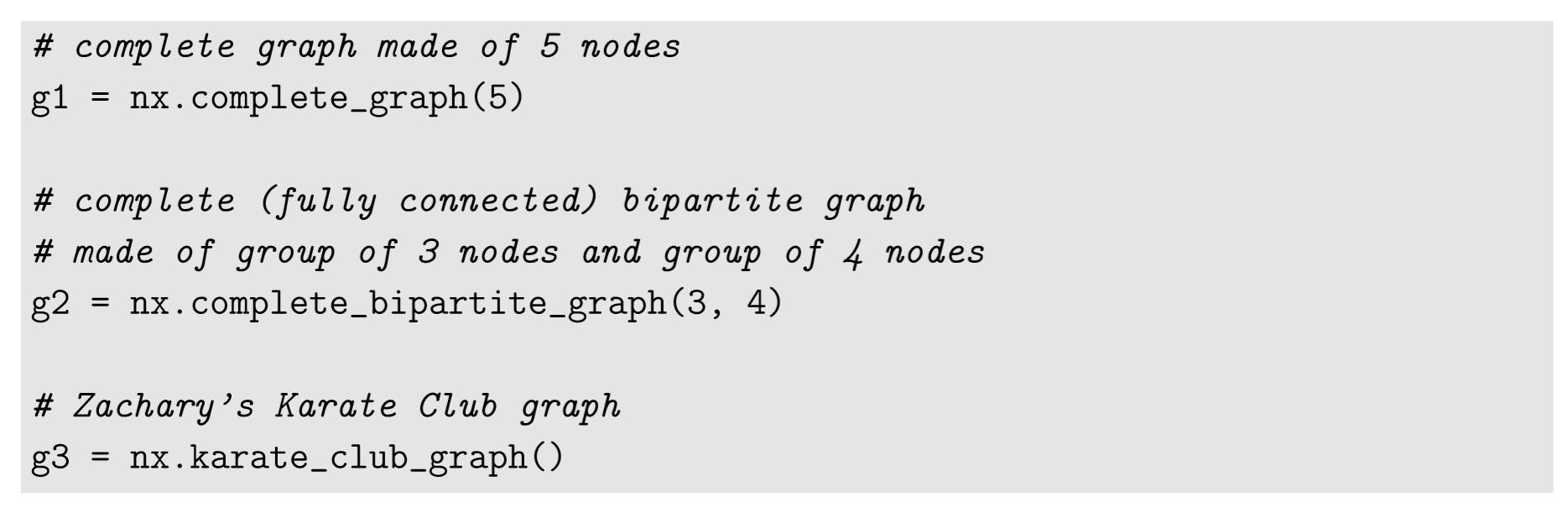


The last example (Zachary’s Karate Club graph) is a famous classic example of social networks reported by Wayne Zachary in the 1970s [59]. It is a network of friendships among 34 members of a karate club at a U.S. university. The edge lists of these examples are as follows:


Construct a graph by generating a complete graph made of 10 nodes, and then connect a new additional node to each of the 10 nodes, using one edge each.
Create Zachary’s Karate Club graph using NetworkX’s built-in function, and inspect its nodes and edges to see if they have any non-topological properties.


